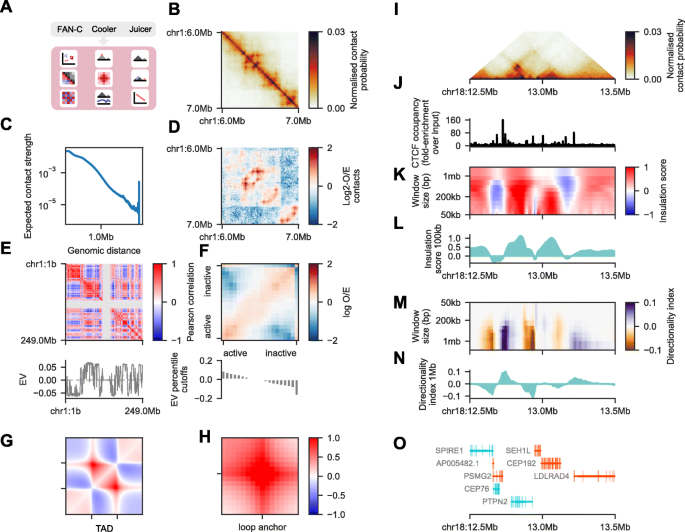
FAN-C: a feature-rich framework for the analysis and visualisation of chromosome conformation capture data | Genome Biology | Full Text
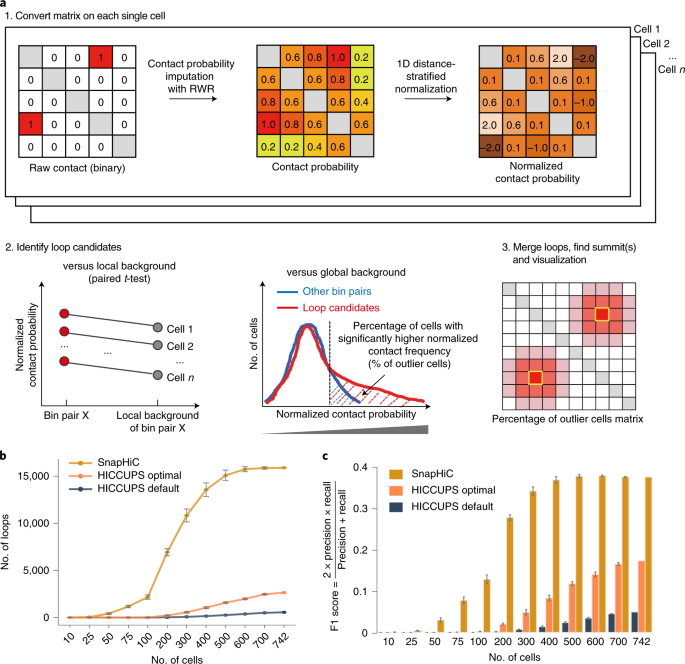
SnapHiC: a computational pipeline to identify chromatin loops from single-cell Hi-C data | Nature Methods
Overview of PHi-C pipeline. Hi-C contact matrix data generated from a... | Download Scientific Diagram

Robin H. van der Weide on Twitter: "Main benefits of GENOVA include: 🗂️ imports Juicer, Cooler and HiC-Pro files 🧰 contains the majority of analyses used in literature 🆕 new tools (!)
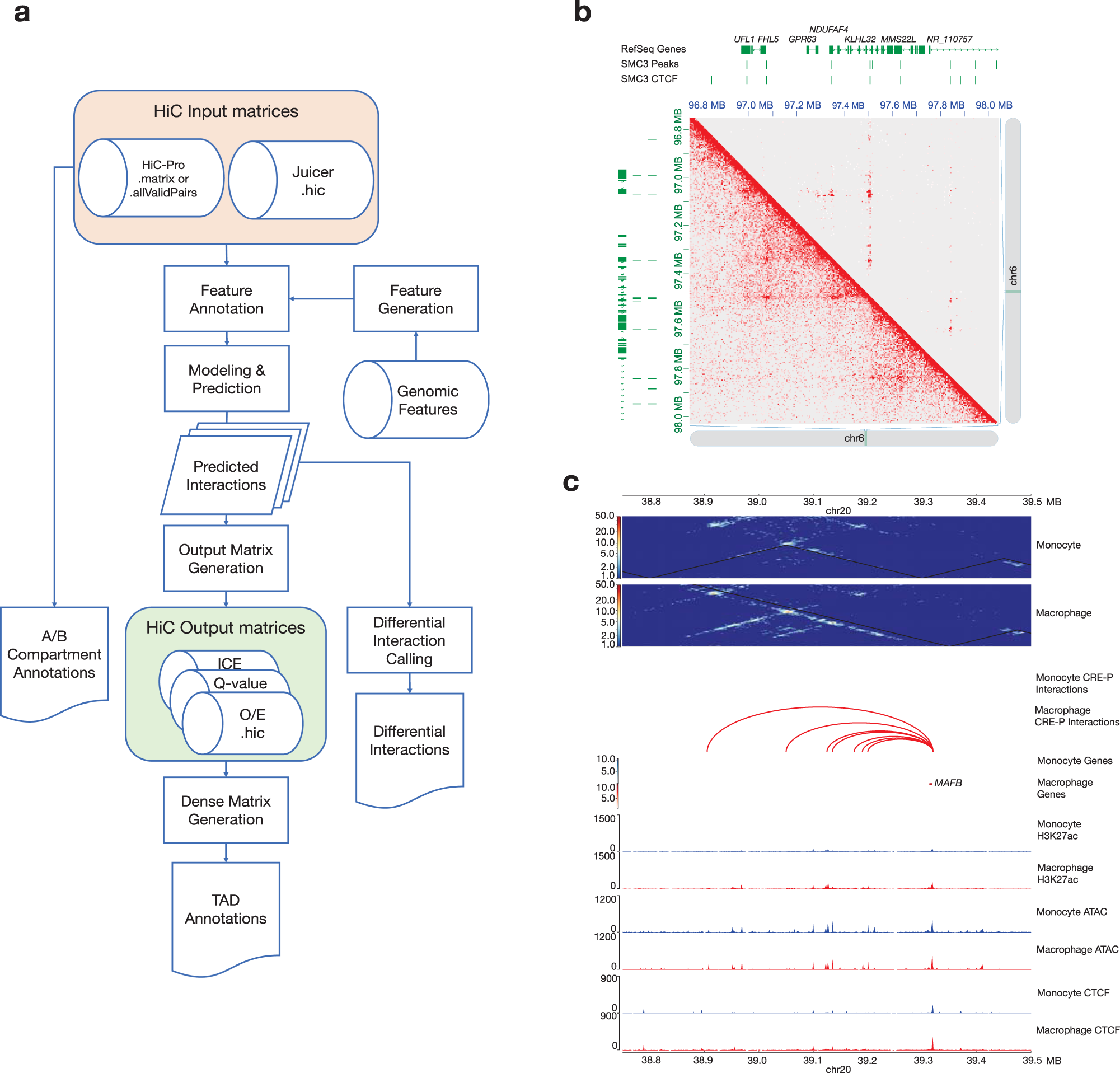
HiC-DC+ enables systematic 3D interaction calls and differential analysis for Hi-C and HiChIP | Nature Communications